Dr. Janneke Ouwerkerk, microbiome and fermentation expert, and Dr. Andrei Prodan, bioinformatics and data science expert, both from the food and health research company NIZO, discuss some of the key issues and challenges facing the nutraceutical and food ingredient industry today.
How do probiotics work to promote health, inside and outside the gut?
Janneke Ouwerkerk: The probiotic can have a systemic effect – i.e., it improves gut health, which impacts overall health – or a more direct effect outside the gut. For example, certain probiotics create butyrate, a short-chain fatty acid that offers many possible health benefits. The butyrate flows through the gut barrier and mucus layer and provides energy to the epithelial cells, thereby strengthening the intestinal barrier function.
Bacteria can also improve our ability to digest food and get more energy and nutrients out of it. Some bacteria can help improve our immune systems, or produce beneficial compounds such as fatty acids or even bacteriocins that kill off ‘bad’ bacteria. In a more general way, by colonising the niche in the gut with ‘good bacteria’, we may prevent pathogenic bacteria from gaining a foothold.
Bacteria are diverse living organisms, so there isn’t a single mechanism for all of them. Even within bacteria species, different strains can have different mechanisms of action.
What are ‘traditional’ versus ‘next-generation’ probiotics?
J.O.: The ‘classic’ bacteria familiar to the industry include Lactobacilli, Bifidobacteria, Streptococci, Saccheromyces (baker’s yeast) and Escherichia coli. But increasingly, we are seeing the use of more novel, or next-generation, bacteria. Faecalibacterium prausnitszii, Akkermansia muciniphila, Anaerobutyricum soehngenii and Bacteroides and Roseburia strains are a few.
These next-generation strains are naturally part of the human microbiome, so they are much more likely to successfully colonise the gut, or be metabolically active. Furthermore, using bioinformatics, they can be targeted scientifically for specific health benefits. But they can also be more difficult to work with, and we know less about them and their mechanisms of action.
What are the steps to choosing and developing a probiotic?
JO: The first step is to identify which strains you want to explore. This discovery starts in silico. A lot of information is already available, which can be used to predict a strain’s behaviour and probiotic potential, but you have to know what you are looking for. Bioinformatics, for instance, can be a powerful screening tool.
Once an initial selection is made, the bacteria have to be produced in large quantities. However, they can be quite difficult to grow. Many next-generation bacteria are strict anaerobes from the zero-oxygen environment of the gut. You need to develop an optimised process that includes appropriate growth conditions.
Then you can begin upscaling, to test the production process, but also to produce enough bacteria for stability and in vitro trials. Step by step upscaling is critical, because even small changes can have a big impact on how a bacterium reacts. The in vitro phase further screens potential bacteria, before clinical tests to validate health benefits.
What challenges must the manufacturer keep in mind?
JO: One of the main challenges is ensuring sufficient probiotic yield and stability. In high throughput screenings, you can already begin eliminating bacteria that grow too slowly, die off too quickly, or do not meet the requirements for good manufacturing practices and food grade production. In silico genome safety assessments can also already reveal antimicrobial resistance genes, toxigenic genes or pathogenicity islands that might remove the bacterial strain from consideration.
By considering aspects such as the transfer to the contract manufacturing organisation and intellectual property (IP) rights early on, you can avoid bottlenecks later.
How can bioinformatics make the probiotics journey more successful?
Andrei Prodan: During the in silico phase, bioinformatics can screen hundreds or thousands of bacterial strains simultaneously. Sequenced genomes can be used to weed out bacteria with dangerous traits, such as antibiotic resistance or virulence. Remaining strains can then be screened for genes linked to targeted health benefits in the literature.
Bioinformatics can also provide insights that are useful in later clinical trial design. For instance, whether a strain could potentially produce certain fatty acids or metabolise a specific substrate in the gut.
The data science facet of bioinformatics helps to obtain useful understanding from in vitro experiments. Data from different equipment and assays, for dozens of strains and tested media, are consolidated and synthesised into expressive visualisations. Endless rows of data are distilled into clear, meaningful insights that can be used to decide which strains or culturing conditions to pursue.
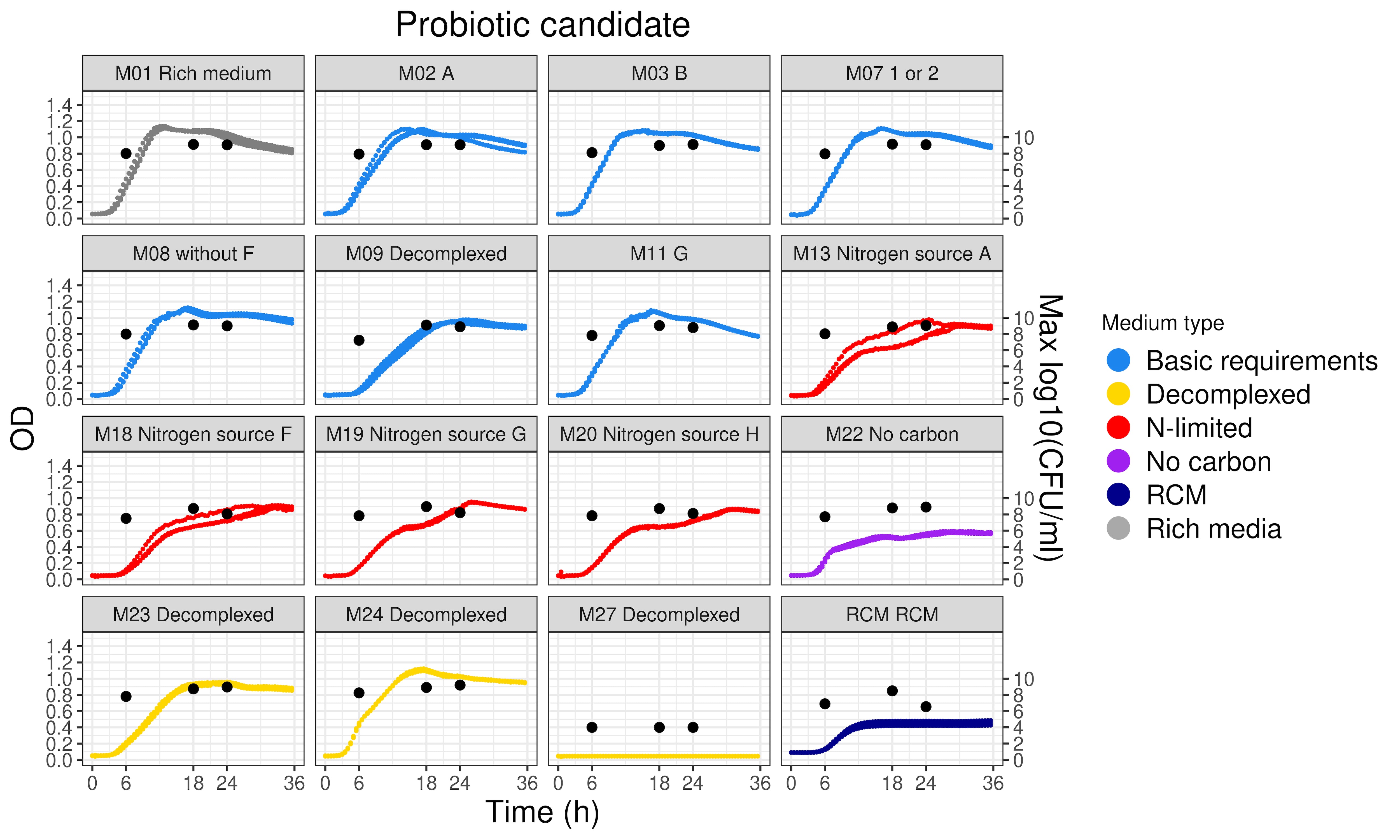
In addition to reducing e.g., 400-500 strains down to 20-40 for testing, in silico screening-based ‘strain passport’ can be made listing potential health benefits ranging from metabolic balance to mental health, sports benefits, sleep-related benefits, and more.
It’s important, however, that bioinformatics be part of a multidisciplinary approach, combined with, for example, microbiology and a technical knowledge of the lab equipment.
What are some of the upcoming trends in probiotics?
J.O: Probiotics will be found in many more different types of products: fruit drinks, energy bars, breakfast cereals…. And they will be targeted at more diverse health benefits: skin, vaginal or mental health, etc.
We will also see more multi-strain products, containing more than one probiotic, to strengthen the specific health benefit. Symbiotics, which combine pre- and probiotics or pro-and postbiotics, are another up-and-coming trend.
Overall, there is a greater awareness of the body’s different microbiomes, and this is going to drive interest in probiotics in new ways.