Just as a person’s blood type comes in one of eight types, research published Nature suggests that everybody has a gut microbiome which falls into one of three clearly distinguishable populations types.
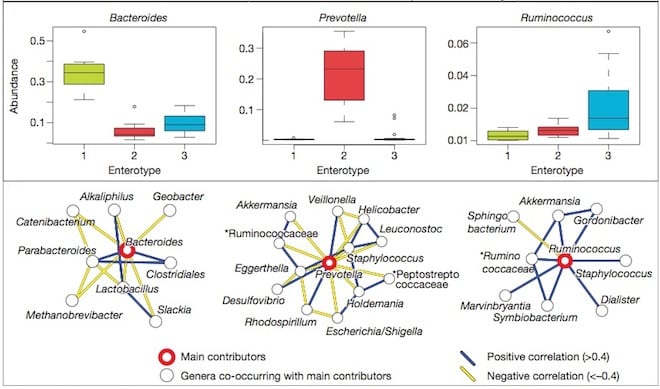
The metagenomic study from a global team of scientists, found that these types are not related to race, native country or diet. The researchers said that types of gut microbiota (called enterotypes) can be classified into three large, clearly distinguishable groups: Bacteroides, Prevotella and Ruminococcus – so names after the main bacteria to dominate the guts of the respective groups.
“The three gut types can explain why the uptake of medicines and nutrients varies from person to person,” said Jeroen Raes a bioinformatician at VIB and Vrije Universiteit Brussel – one of the two lead researchers in the study.
The researchers added that their findings may have major implications for detecting and predicting risks of disease such as intestinal cancer, diabetes, and Crohn's disease.
Commenting independently on the research, prebiotics pioneer Professor Glenn Gibson from the University of Reading told NutraIngredients that the value of the new findings is that the high throughput approach used to model the data and the direction “towards genus/species level information” on the understanding of gut microbes..
He added that it makes sense to try and understand the gut microbiome at a species level, “as dietary intervention strategies with prebiotics and probiotics act at those levels.”
“The target microbes for such approaches should be understood,” said Gibson
Gut ecosystem
Although we have known for hundreds of years that we have bacteria living in our guts, it is only recently that new technologies have allowed researchers to begin to understand the size and complexity of the ecosystem living within us.
An estimated 100,000 billion individual bacteria live in our intestines, making up the complex ecosystem that is the gut microbiota. These bacteria help to break down food, protect us against attacks by pathogens, and play a vital role in health and nutrition.
In a recent interview with NutraIngredients, Professor Tine Rask Licht, an expert in gut health, and the winner of the 2010 Danisco Award for her research in the field, said that recently there has been a shift towards starting to consider the gut microbiota as an organ in its self.
In her exclusive interview, she added that the knowledge and indications to suggest that gut microflora interact with the risk factors for lifestyle associated diseases and human health has “really exploded” in recent years.
The metagenomic technique studied the genetic information of all the bacteria in present in faeces, generating a vast amount of data that is then analysed for patterns.
Study details
The research team combined 22 newly sequenced faecal metagenomes of individuals from four countries with previously published data sets; finding that three robust clusters that are not nation or continent specific.
Raes and his colleagues reported that although individual host properties, such as body mass index, age, or gender did not explain the observed enterotypes, other data markers such as genes and functional modules can be identified for each of these host properties.
“For example, twelve genes significantly correlate with age and three functional modules with the body mass index, hinting at a diagnostic potential of microbial markers,” they explained.
The researchers also found that vitamin production between the different groups varied strongly.
They noted that people that belong to the Bacteroides group have more gut bacteria that produce vitamins C, B2, B5 and H, whereas thePrevotella group showed higher numbers of B1 and folic acid-producing flora.
Implications
The researchers explained that they do not yet have a conclusive explanation for the existence of the types of gut microbiota, but said that all the evidence indicates that only a limited number of stable bacterial communities are possible.
Raes and his co-workers added that the enterotypes appear complex, and “are probably not driven by nutritional habits.”
They also warned that whilst the currently identified enterotypes were ‘robust’, their findings are based on a relatively small sample of people, meaning further research is needed to research determine whether there are additional, as yet unidentified, gut bacteria population types.
Source: Nature
Published online ahead of print, doi: 10.1038/nature09944
“Enterotypes of the human gut microbiome”
Authors: M. Arumugam, J. Raes, E. Pelletier, D. Le Paslier, et al
