Called the AiMS (automated in vitro microbiome screening), the system was developed by a team led by Andrew Benson, PhD, head of the center, which is part of the University of Nebraska-Lincoln’s Department of Food Science and Technology.
Like many stories in the development of new ingredients, the development of AiMS was an overnight success that took years.
“We’ve been at this for more than six years, and four of those were spent just in optimizing the process,” Benson told NutraIngredients-USA.
Trying to replicate a complex system
The hurdles in the research of the effects of ingredients on the gut are many, Benson said. A big one is the sheer complexity, both in terms of upstream reactions (what happens in the stomach and upper parts of the small intestine) and the vast, still only dimly understood world of the colonic microbiome itself.
“Trying recreate that exact ecosystem is essentially impossible,” Benson said. “The best you can do is to put in the components you know should be there and then measure the interactions. And from there you have to be able to extrapolate.”
Developing in vitro models of digestion is nothing new, Benson said. The real problem is scaling a model in such a way that really significant amounts of data can be gathered, improving the statistical power.
Making it fast, easy and reliable
Benson said the problems to be solved were similar to the mass production of almost anything: reproducibility and cost. Having humans (even of the eager grad student type) tending individual test tubes or petri dishes is time consuming, expensive and rife with potential error. Benson said the team had to find a way to scale that process so that it was quick, cheap and reliable.
“We took an existing system that had been validated in 5 ml tubes and scaled it to use on 96 well plates,” he said.
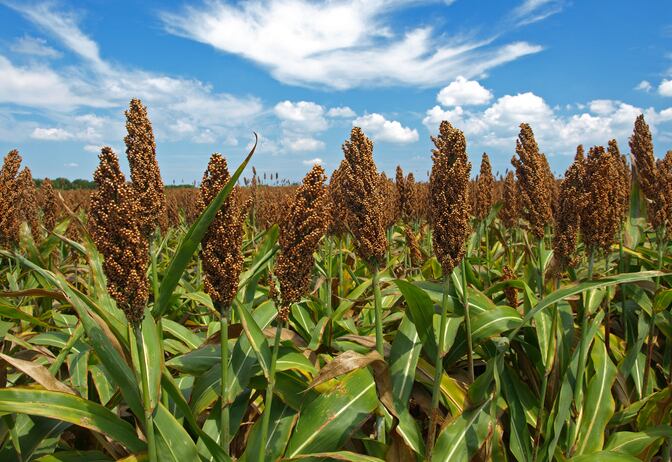
The system, which was a key feature of the team’s recently published research on prebiotic potential in high tannin sorghum varieties, relies on some key equipment choices. One was a special grinder unit sold by ThermoFisher Scientific that most researchers use to prepare samples for DNA extraction, Benson said. In his team’s case, the unit was used to do precise milling of hundreds samples for distribution into the test tubes.
Distributing the samples
Distribution of the test materials with the next hurdle, Benson said. What the Nebraska team wanted to do was somewhat unique in a world of laboratory materials distribution.
“There are systems out there that can distribute from many into one, like into a machine to make capsules or tablets,” Benson said. “But in our case we wanted to distribute from many into many.”
The team finally found a piece of equipment from Swiss firm Chemspeed that features a robotic arm that could precisely dose the test material into the tubes. Having grad students, however willing, dispense material over and over is the kind of mind numbing task that humans don’t do well but where properly calibrated machines excell, Benson said.
“Trying to weigh out 20 mg a thousand times over and over—you wouldn’t have too many grad students stick around with you for that,” Benson said.
Simulating stomach action
That material then needed to be treated to resemble its condition after having already gone through digestion in the upper GI tract, which included treatment with acids and enzymes. It then needed to be gently stirred so that those digestion products would pass through the filter in the bottom of the plate that would simulate absorption through the intestinal lining, leaving behind the indigestible fractions that were the subject of the study.
The stirring was accomplished with tiny magnetic beadlets dropped into the tubes. These are activated by a magnet rotating around the water-filled buckets in which the plates were suspended. That system was another key technical piece of the puzzle, Benson said.
Distributing the stool samples
The Chemspeed unit was also used to distribute the cultured microbiome samples, which in the case of the sorghum study came from stool samples taken from 12 individual donors.
“When I described our needs to them, the Chemspeed people said, ‘You want to do WHAT with our machine?!” Benson joked.
Getting enough of the microbiome samples to distribute into thousands of different tubes is a less onerous and nasty job than it used to be, Benson said. A team leader like himself can only prevail so far on his graduate assistants. Systems used in clinical setting exist for the—relatively—painless proliferation of stool samples.
Benson said in the end the team has a validated system that holds great promise for future research. The team met most of the goals it set out to achieve. One thing that’s hard to get around is the high number of pricey consumables inherent in the approach. The cost of many of those rose rapidly during the pandemic.
“It’s fast and reliable, yes. Cheap? I’m not so sure,” Benson said.